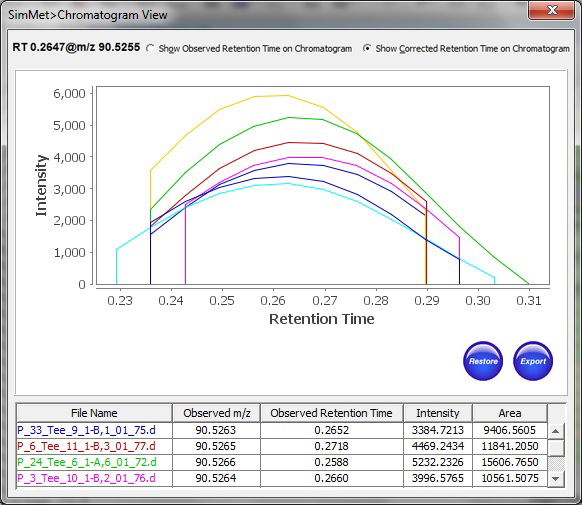
Mestrelab Research on X: "Interested in LC-MS data analysis for Open Access? Our colleague Dr. Camil Joubran described how to process, analyze and report your LC-MS data from different instruments using Mnova
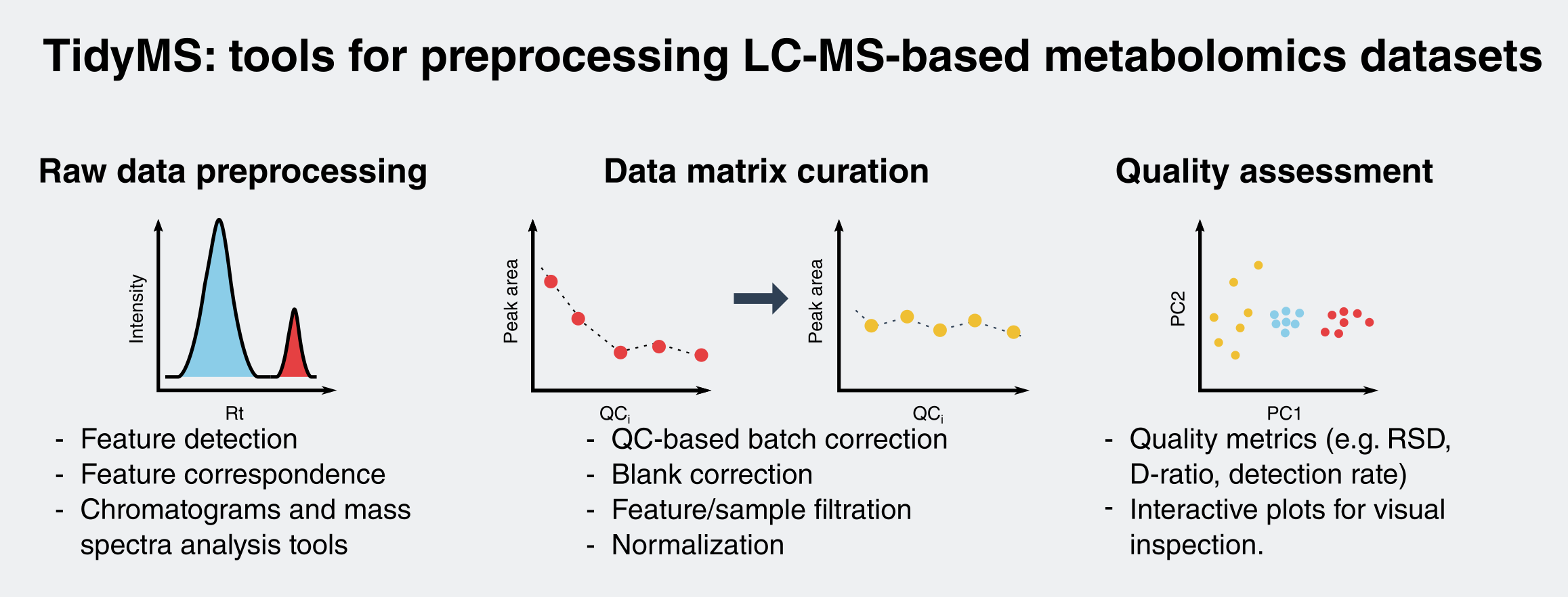
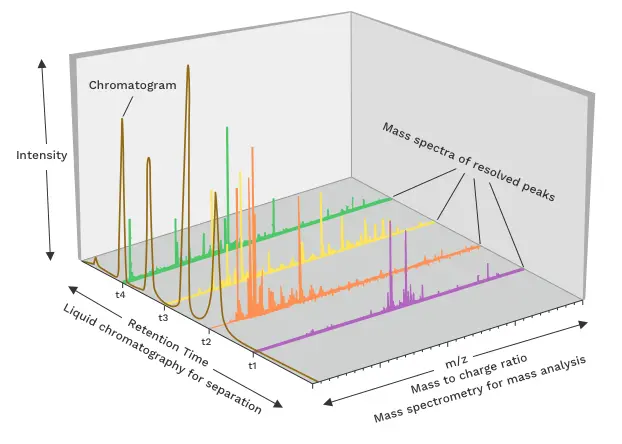
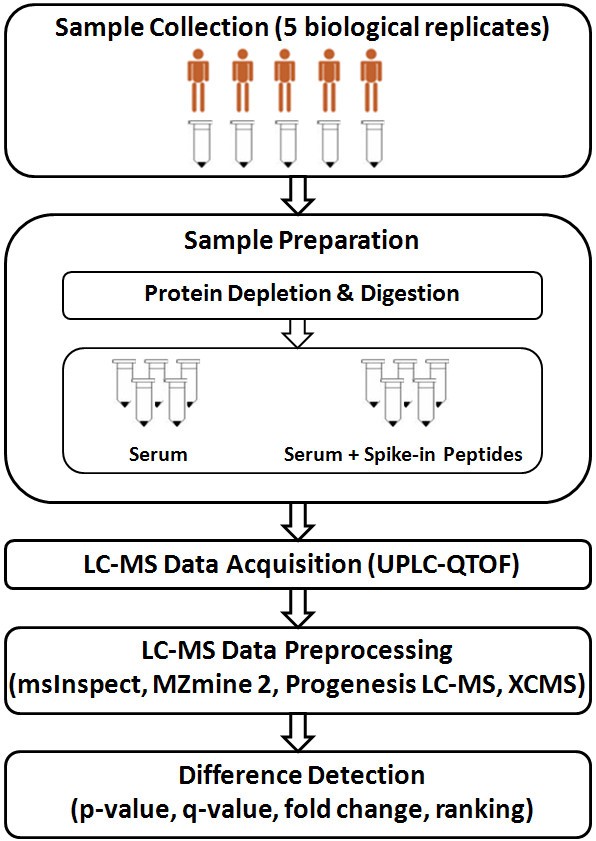
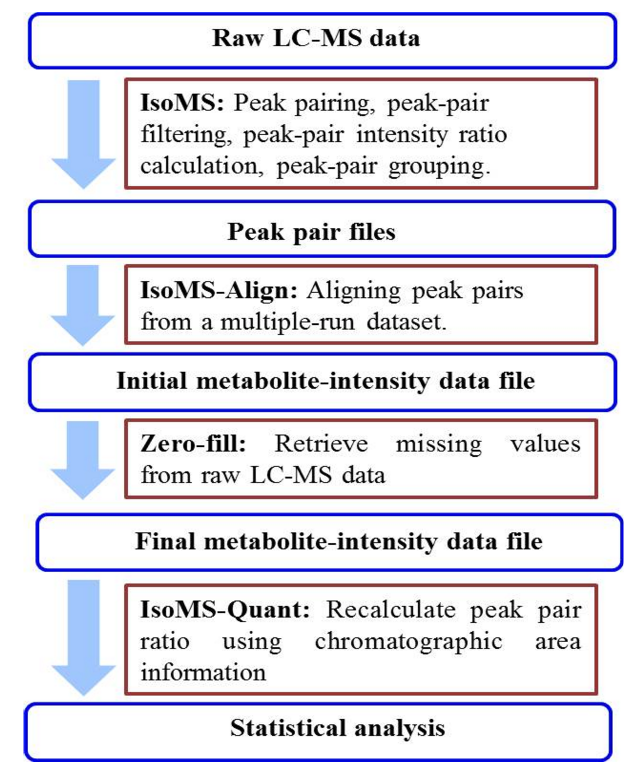
Metabolites | Free Full-Text | A Python-Based Pipeline for Preprocessing LC–MS Data for Untargeted Metabolomics Workflows

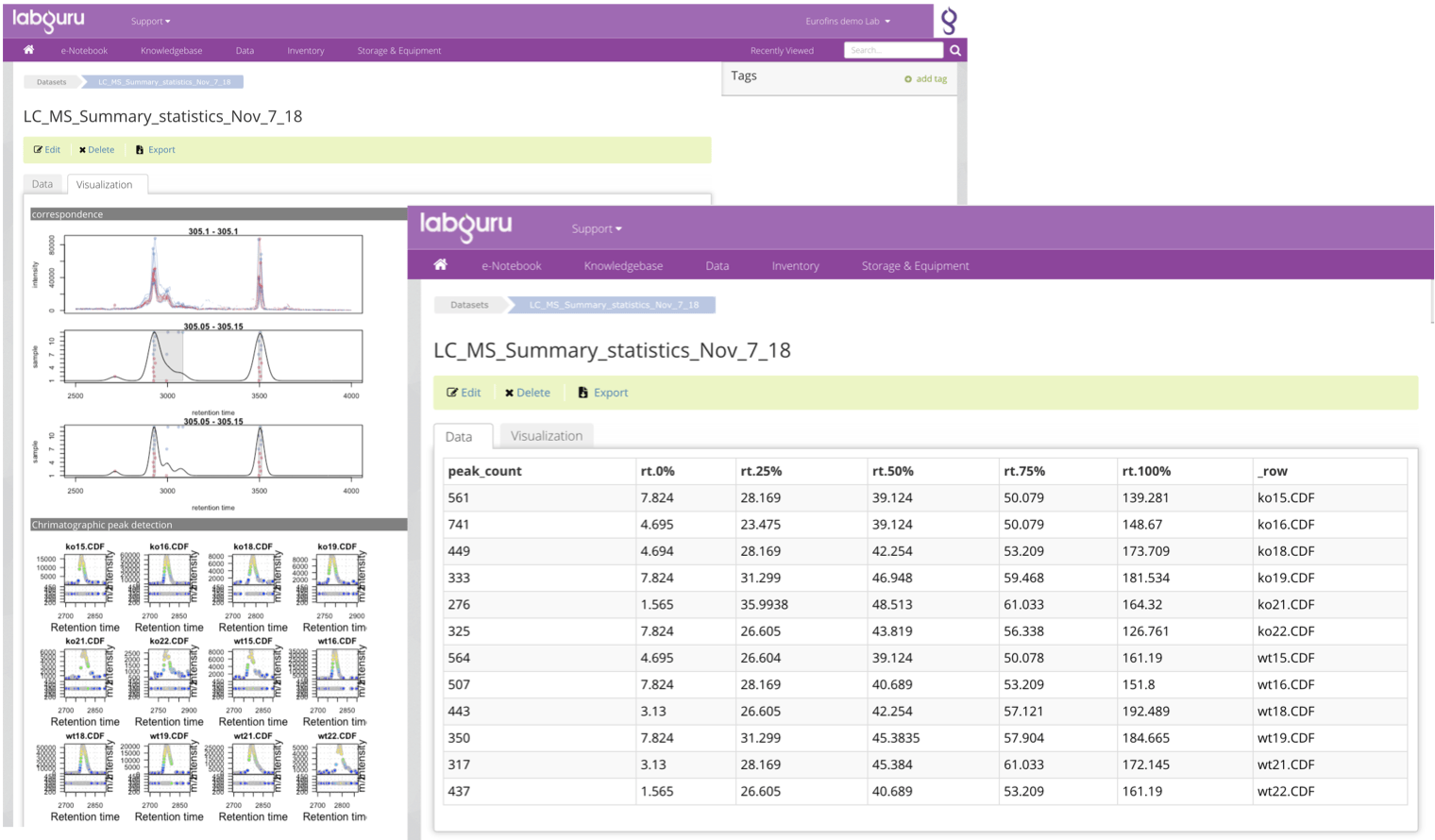
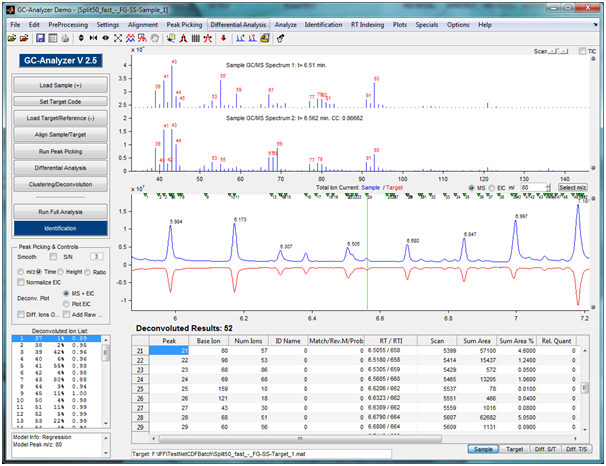
Improving LC/MS Workflows efficiencies with Batch queue Sample Submission and Automated Data Analysis

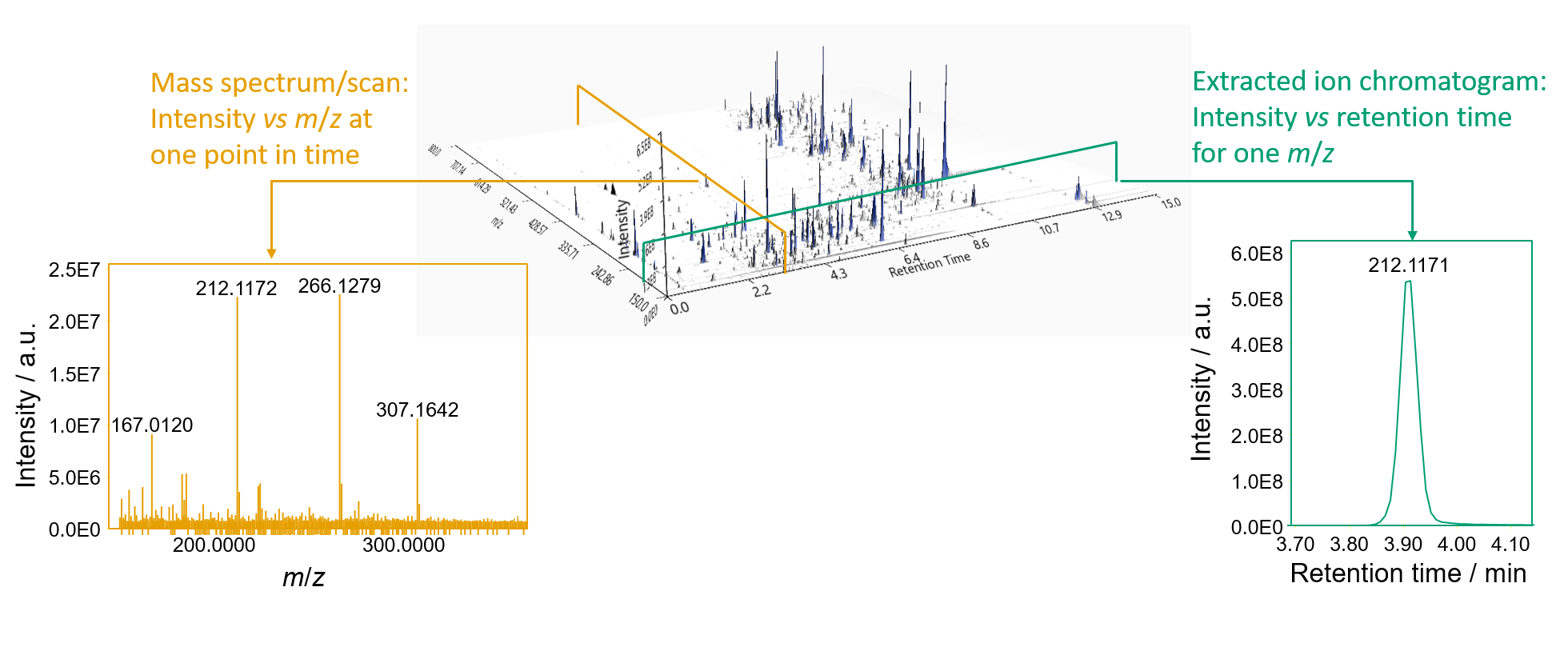
LC MS and MS/MS data from whole blood. a MS data from an LC fraction... | Download Scientific Diagram
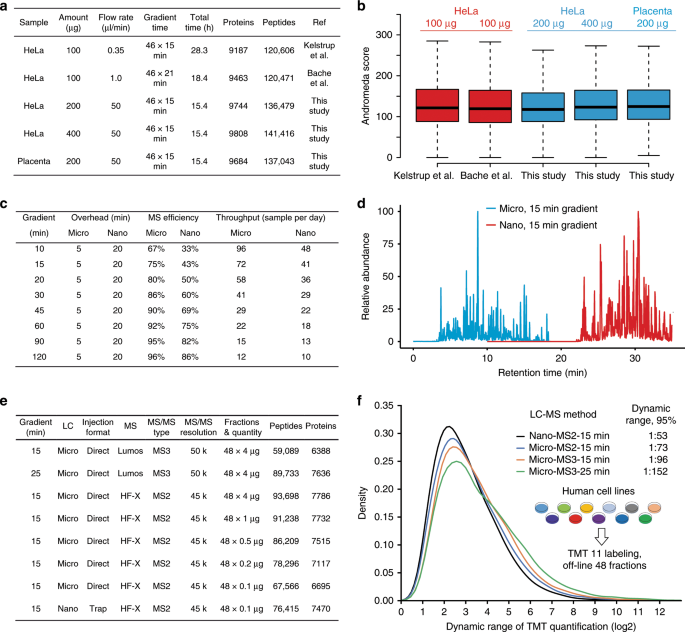
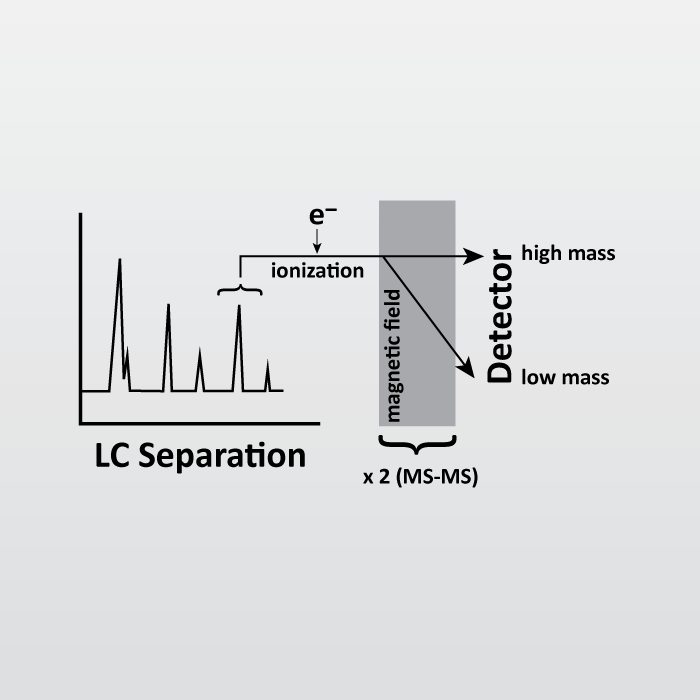
Robust, reproducible and quantitative analysis of thousands of proteomes by micro-flow LC–MS/MS | Nature Communications
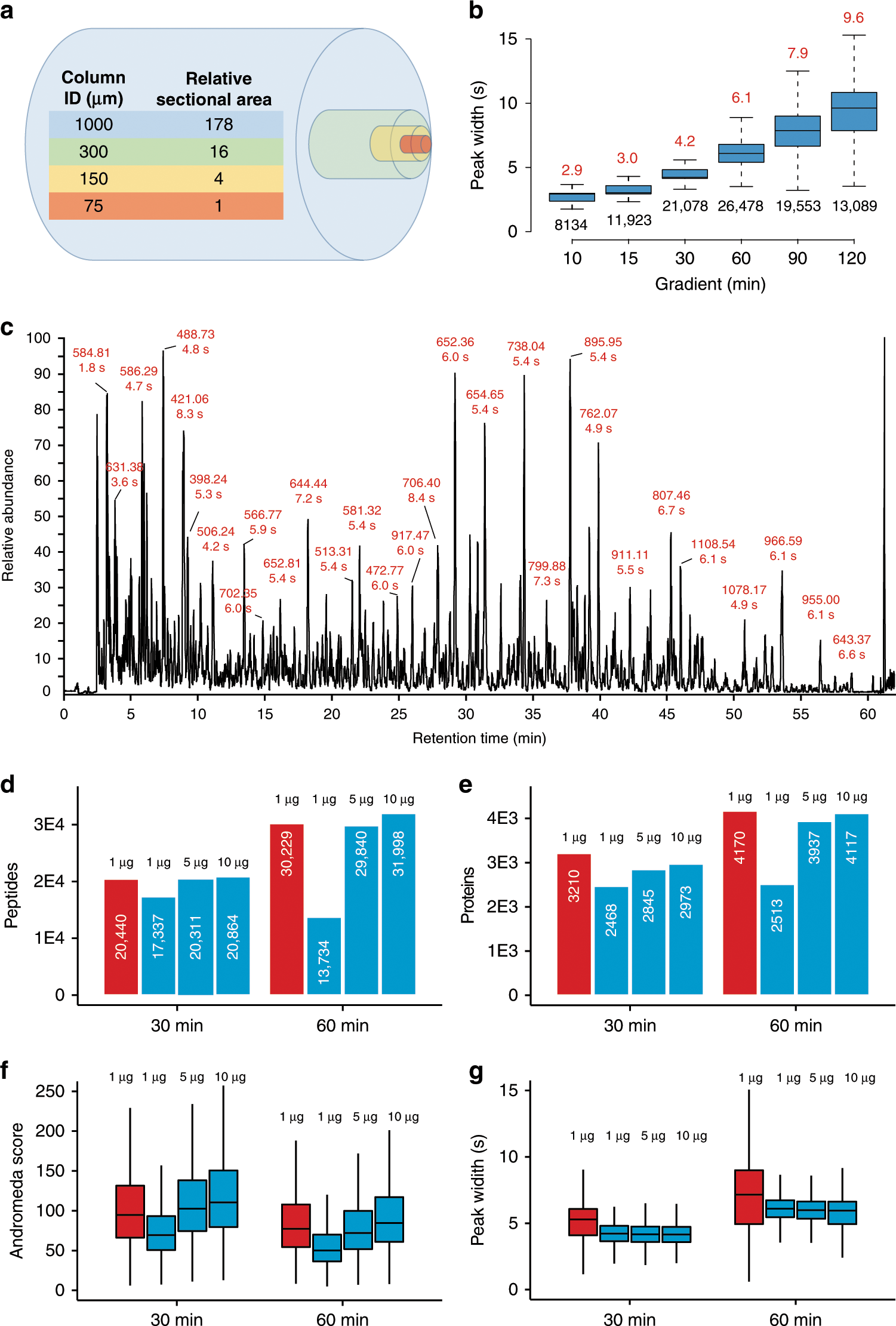
Robust, reproducible and quantitative analysis of thousands of proteomes by micro-flow LC–MS/MS | Nature Communications

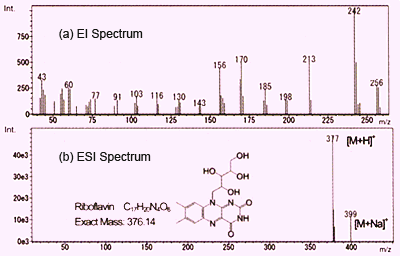
Small Molecule LC-MS/MS Fragmentation Data Analysis and Application to Siderophore Identification | IntechOpen